
The Alcoholic Hepatitis Network project, AlcHepNet (http://www.alchepnet.org/), is sponsored by the National Institute on Alcohol Abuse and Alcoholism (NIAAA).
AlcHepNet aims to improve the treatment of alcohol-associated hepatitis, a leading cause of liver-related morbidity and mortality. The AlcHepNet consortium comprises eight clinical study sites and ten research projects, including clinical, translational, and primary or preclinical studies. The network is recruiting over 1,300 participants for clinical studies, following up with participants for 180 days, collecting more than 78,000 blood, urine, saliva, and liver biopsy bio-samples, capturing demographic and behavioral features, clinical conditions, laboratory tests, treatments, and outcomes, and generating multiomics data from microbiome, immunologic, proteomic, metabolomic, lipidomic, and RNA/ChIP-sequencing analyses. The Indiana University Data Coordinating Center (IU DCC) and the University of Massachusetts Data Coordinating Center (UMass DCC) are collaboratively providing the essential research infrastructure, including experimental design, study implementation, data management, and statistical analysis in support of the two primary studies within the network, a clinical trial and an observational study, as well as the translational projects that utilize the biospecimens collected by the two primary studies. To facilitate effective research use of the rich and complex AlcHepNet data, the IU DCC has developed ARDaC , the Alcohol Research Data Commons, as the central data hub and research nexus (https://portal.ardac.org/).
... Read More
Design of ARDaC
The architecture of the ARDaC system is composed of the following components:
-
The ARDaC Data Warehouse. The heterogeneous clinical data, biosample information, and omics data information will be extracted from the IU DCC and UMass DCC,
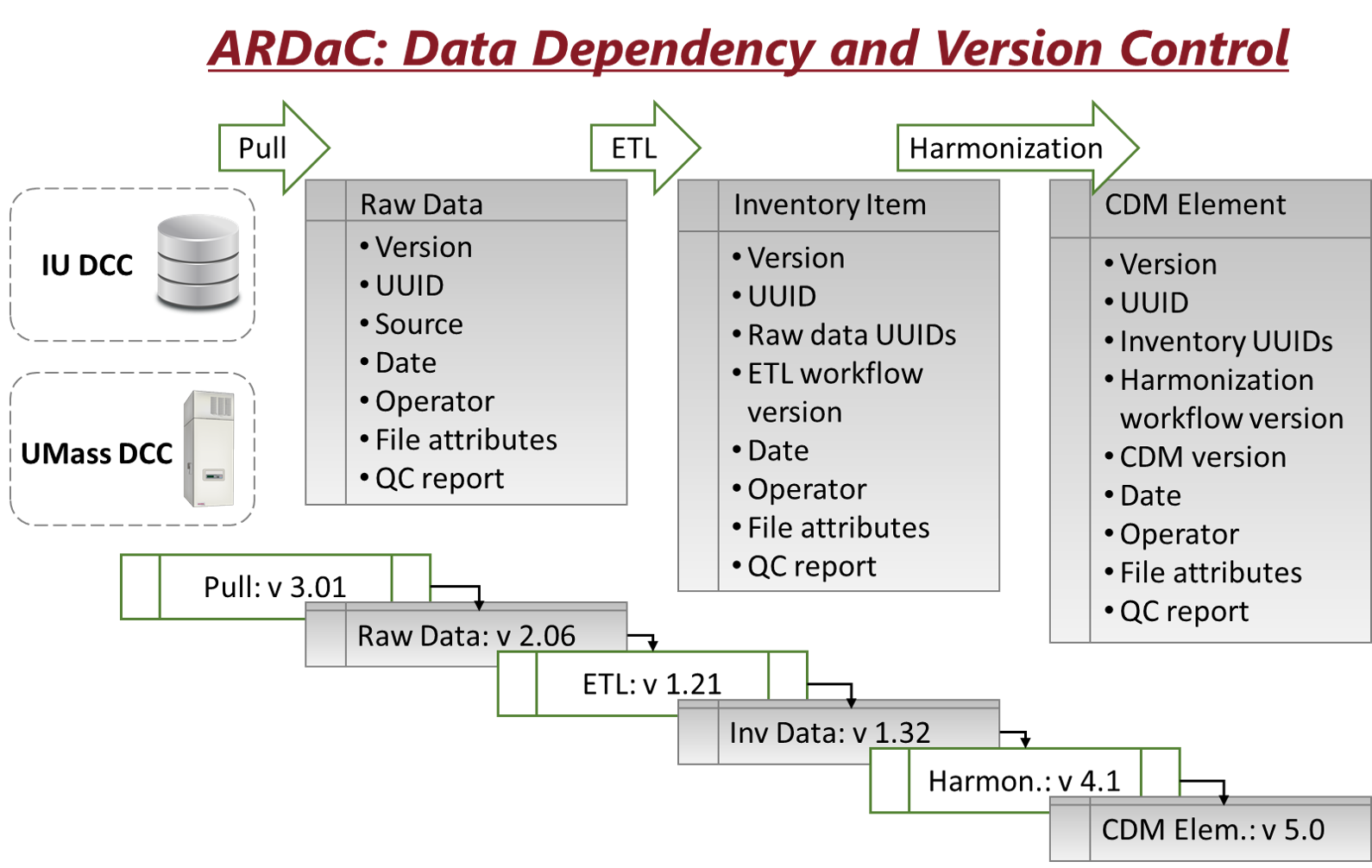
standardized according to the ARDaC Data Standard, harmonized according to the ARDaC Common Data Model, and hosted in a central ARDaC Data Warehouse. Specifically, the novel ARDaC Common Data Model is derived from and compatible with the Genomics Data Common (GDC)[1] Data Model and is compliant with the FAIR Principles[2] so that AlcHepNet multimodal data will be findable, accessible, interoperable, and reusable. The ARDaC Data Warehouse is the data source for the ARDaC web application, which is open to the public, as well as for regular reporting and customized services within the AlcHepNet consortium. A graph-based provenance model is used for comprehensive data dependency and version control. The ARDaC digital entities, including the standards, data model, data, metadata, scripts, and codes, are attributable, trackable, and reproducible.
... Read More
-
The ARDaC web application. The ARDaC system uses the Gen3 data common framework, which is widely used in NIH-sponsored projects.
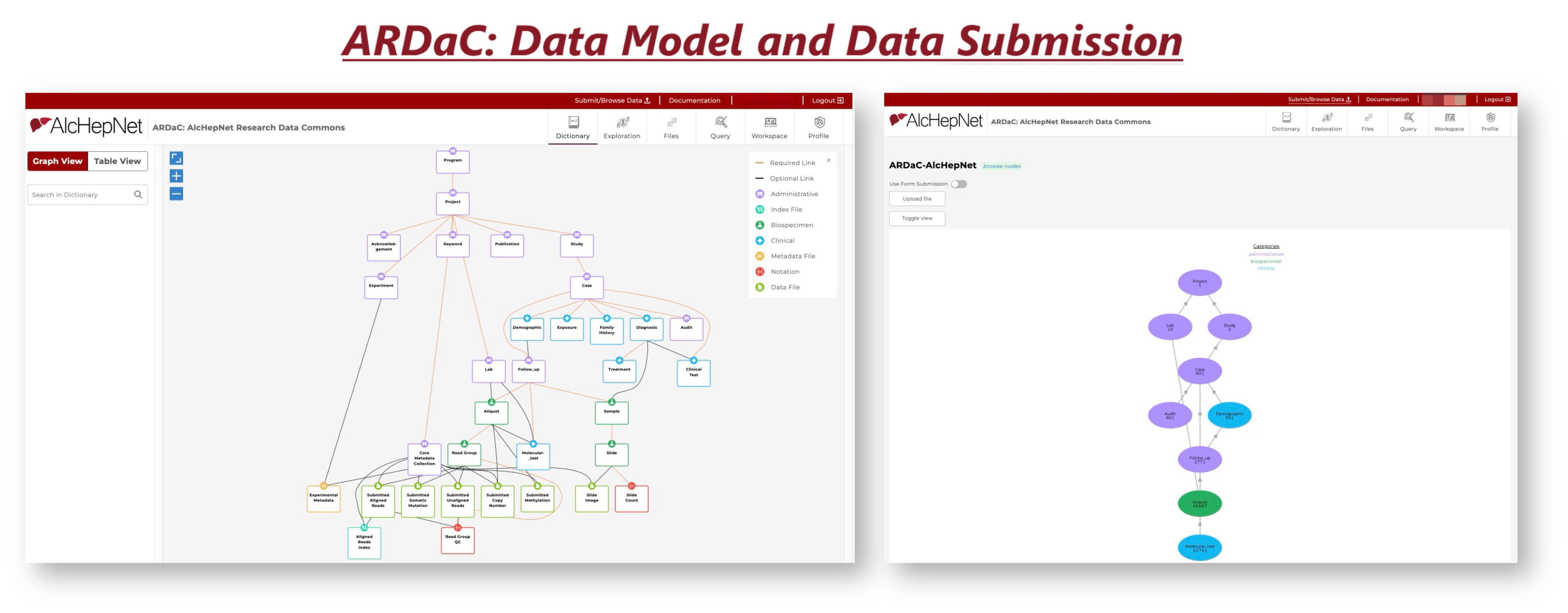
At the data layer, the standardized and harmonized data is extracted from the ARDaC Data Warehouse and injected into the ARDaC Staging Data Warehouse according to the ARDaC Graph Data Model. In the middleware layer, based on the user's input of the filtering criteria, the graph-based data is queried using GraphQL through the elastic search engine, analyzed with Python, and delivered interactively to users using the JavaScript-based react and Storybook libraries. The ARDaC web application is containerized as a series of images, each providing a specific service. The ARDaC system can be deployed to the AWS cloud services through the Kubernetes platform or to a dedicated server through the Docker platform. By leveraging the GDC common data model and the Gen3 data commons ecosystem, ARDaC enables data integration with other NIH-funded data commons, delivering a broad impact of AlcHepNet research and data to other research communities.
... Read More

Functionalities
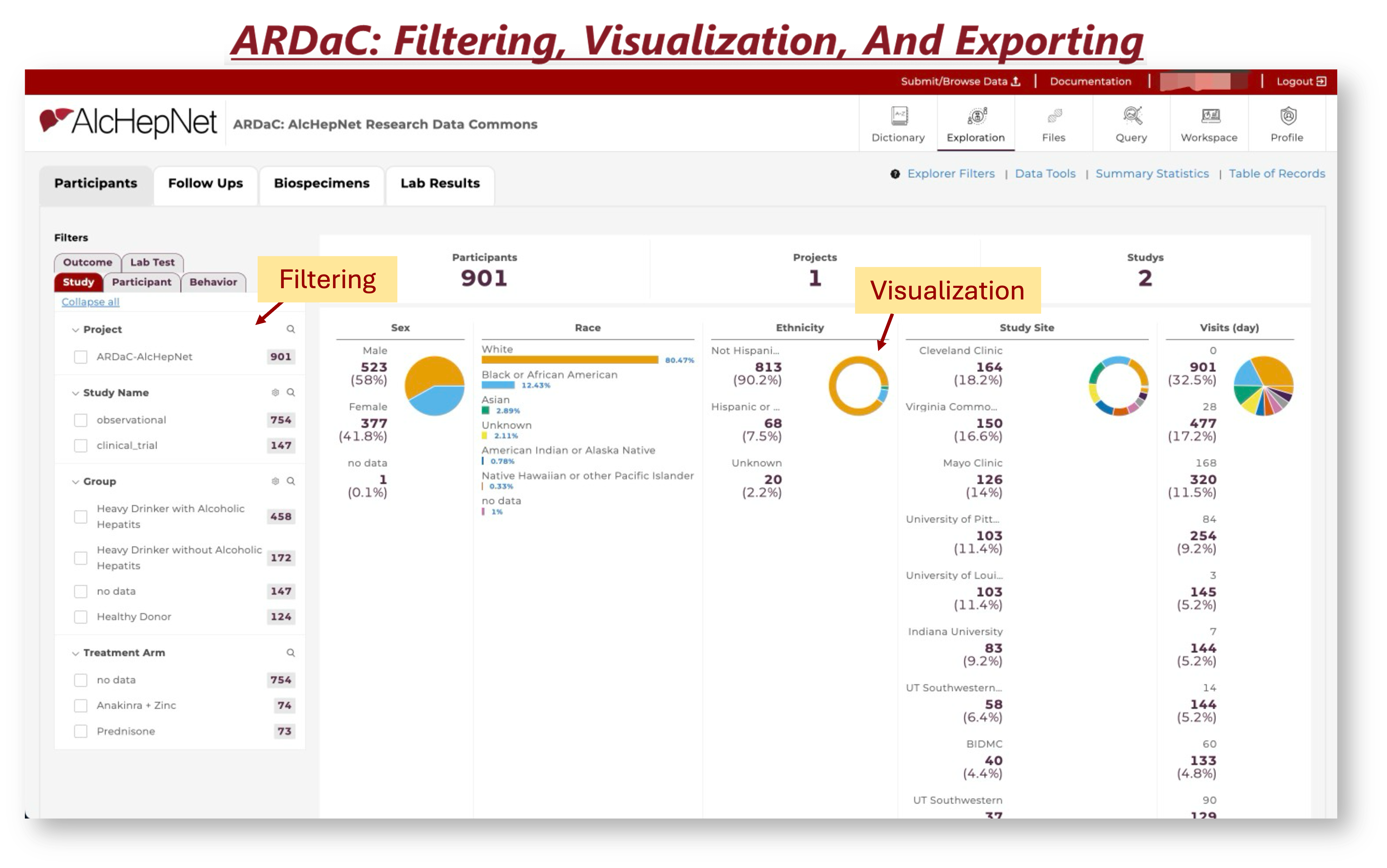
The novel ARDaC system supports the representation of behavioral and pathologic data unique to alcoholic hepatitis and facilitates data filtering, querying, visualization, and exploring,
which are specific to the AlcHepNet clinical studies. Besides the general data query and visualization functions, ARDaC allows data exploration using study-related criteria such as the study cohorts or arms, alcohol use history, alcoholic hepatitis treatments, treatment outcomes such as mortality and liver transplantations, and liver function measures as well as prognostic metrics such as the MELD scores, omics data availability and biosample availability, and omics-derived features such as differentially expressed genes and enriched signaling pathways. The ARDaC system also provides a GraphQL query interface, a cloud-based workspace, and R and Python programming environments for in-depth data analysis. Specifically, ARDaC allows users to check sample availability, visualize and evaluate the synergy of their data generation plans with existing data and funded projects, and plan for new data generation plans. The ARDaC system is available at github.com/jing-su/ardac and ardac.org.
... Read More

In summary, ARDaC is the central data hub connecting data of multiple modalities across clinical and translational teams. It is the engine that drives AlcHepNet research projects, the data interface between the AlcHepNet consortium and other research data commons, and the research nexus that ignites new research and collaborations.
Team
The team at Biostatistics and Health Data Science, Indiana University School of Medicine
PIs:
- Wanzhu Tu, PhD
- Samer Gawrieh, MD
- Jing Su, PhD
Data modeling and harmonization
- Carla Kettler, Principal Data Manager
- Nanxin Jin, PhD Research Assistant
- Ronny Ovando, Senior Data Manager
- Timothy Hotchkiss, Senior Data Manager II
- George Nitsos, Security Analyst
System development
- Nanxin Jin, PhD Research Assistant
Information Technology (IT) support
- Esen Tuna, Research Data Services
- Alan Walsh, Research Data Services
- Guangchen Ruan, Research Data Services
- James McCombs, Research Data Services
- Amy Burns, Enterprise Systems
- Philip Berg, Enterprise Systems
Visualization
- Zuotian Li, PhD Research Assistant
- Hao Wang, PhD Research Assistant
Publications
- N Jin, Z Li, C Kettler, B Yang, W Tu, J Su, "ARDaC Common Data Model Facilitates Data Dissemination and Enables Data Commons for Modern Clinical Studies." Studies in Health Technology and Informatics. 310:3-7(2024). doi: 10.3233/SHTI230916. PMID: 38269754.
https://pubmed.ncbi.nlm.nih.gov/38269754/
- Li, Zuotian, Xiang Liu, Zelei Cheng, Yingjie Chen, Wanzhu Tu, and Jing Su. "TrialView: An AI-powered Visual Analytics System for Temporal Event Data in Clinical Trials." arXiv preprint arXiv:2310.04586 (2023).
https://arxiv.org/abs/2310.04586
Acknowledgements
Contact Us
If you have any questions, please feel free to reach out to us.
Email: ardac@iu.edu


